Overview
This work presents a dynamic template-constrained large language model (LLM) framework for converting free-text lung cancer screening (LCS) radiology reports into fully-structured reports (FSR) with zero formatting errors and zero hallucinations.
Unlike prompt-based approaches, this method constrains every generated token using a predefined clinical template with standardized candidate values. The system was validated across two institutions using 7,442 LDCT reports, achieving state-of-the-art performance.
Clinical Motivation
Radiology reporting traditionally uses loosely-structured or free-text formats (LSR), which limit large-scale statistical analysis and reliable retrieval. Fully-structured reporting (FSR) enables:
- Standardized discrete feature extraction
- Automated statistical mining
- Nodule-level semantic retrieval
- Cross-institutional consistency
- Downstream AI model training
However, conventional LLM prompting fails in clinical settings due to:
- JSON formatting errors
- Content hallucinations
- Privacy risks (cloud-based proprietary models)
- Long inference time and high token cost
📐 Structured Template Design
Two radiologists created a standardized lung nodule template containing 28 features:
- 24 nodule-level features (e.g., lobe, segment, attenuation, margin, shape, size)
- 3 report-level management features
- 1 auxiliary feature (number of nodules)
Each feature has a predefined candidate set (e.g., attenuation ∈ {solid, part-solid, ground-glass, ...}). This predefined schema is the foundation of the constrained decoding.
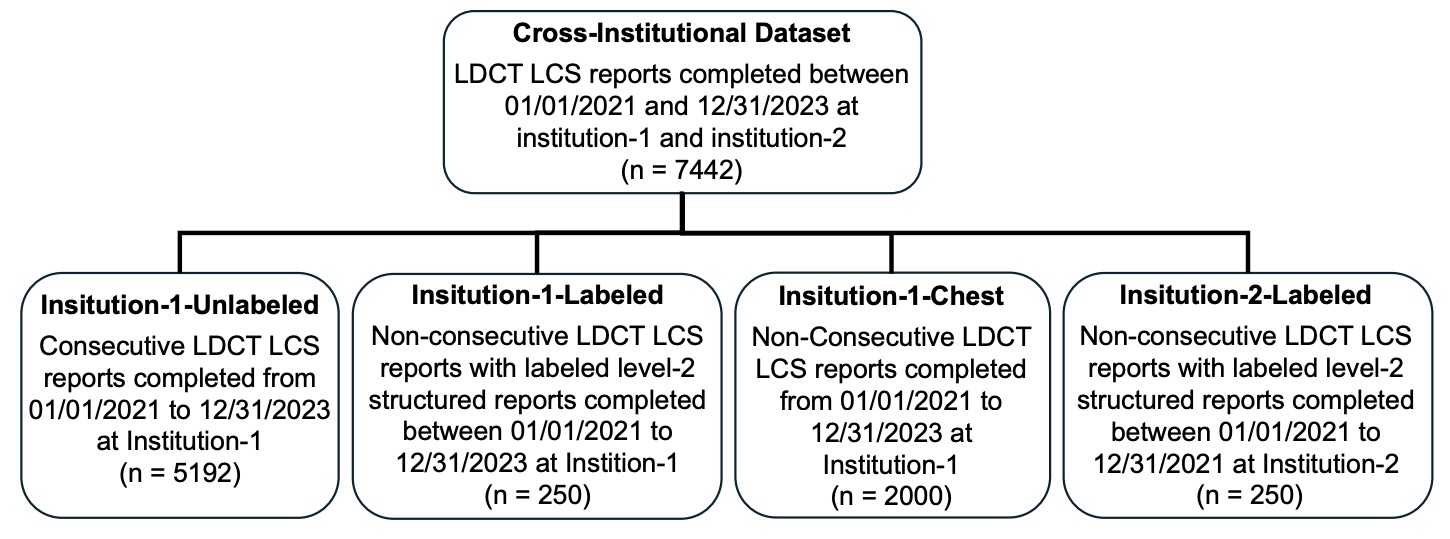
Figure 1: Cross-institutional dataset construction (Institution-1 & Institution-2).
🏗 Dynamic Template-Constrained Decoding
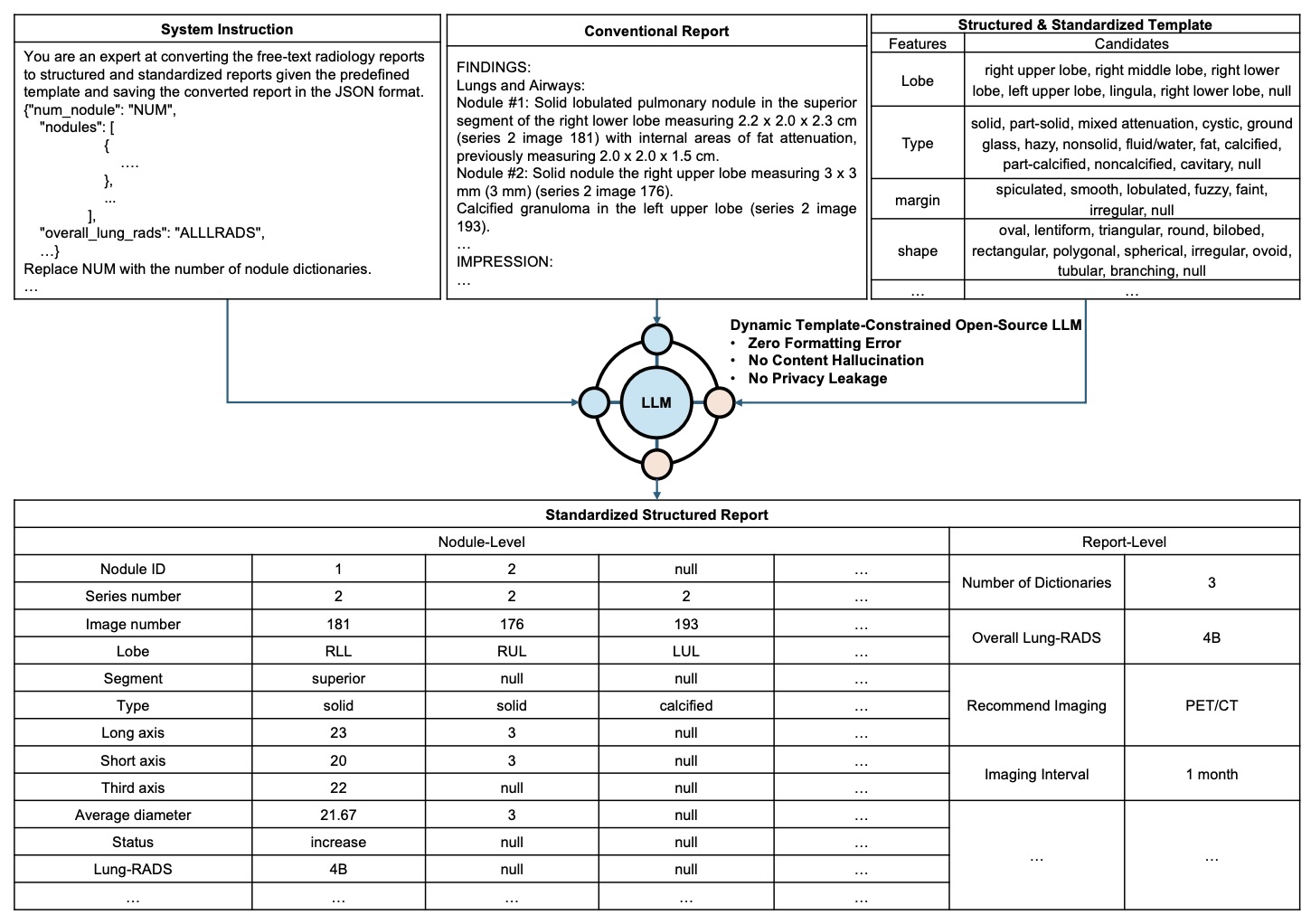
Figure 2: The LLM receives system instruction + free-text report, while decoding is strictly constrained by a structured template.
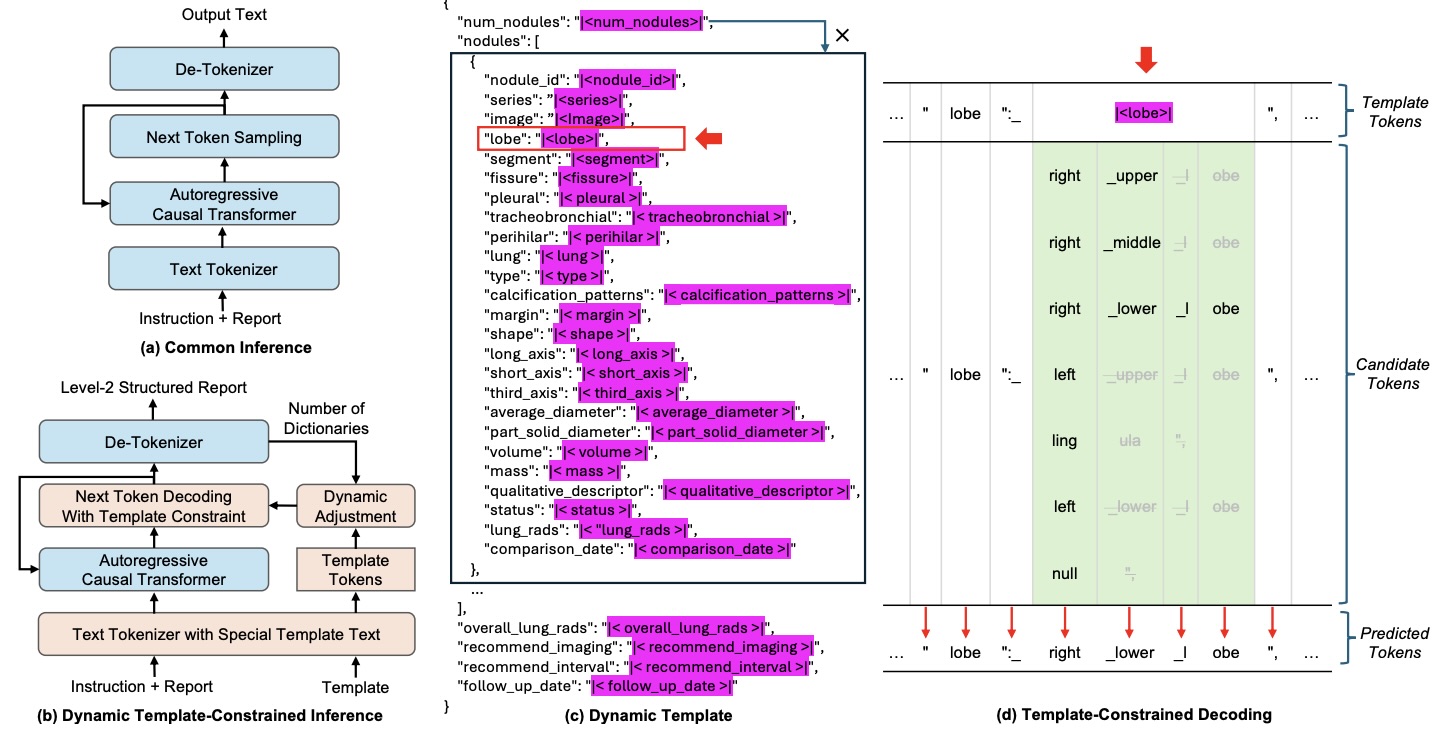
Figure 3: Modified inference architecture integrated into vLLM. Special template tokens restrict candidate outputs during decoding.
During decoding:
- Template format tokens are fixed → ensures valid JSON
- Special template tokens select only predefined candidates
- Dynamic adjustment handles variable number of nodules
- Maximum-probability candidate is deterministically selected
This guarantees:
- 🚫 Zero formatting errors
- 🚫 Zero hallucinations
- 🔒 Local deployment (no privacy leakage)
📊 Cross-Institutional Validation
Evaluation was initially performed on 500 manually labeled reports (250 per institution) to assess generalizability across clinical settings.
- Institution-1: F1 = 97.60% (95% CI: 96.5–98.6%)
- Institution-2: F1 = 96.92% (95% CI: 95.8–98.0%)
Performance Improvements over Baselines:
- LLaMA-3.1-405B: +10.42%
- GPT-4o: +17.18%
- Consistent gains across all 27 nodule features (p < 0.01)
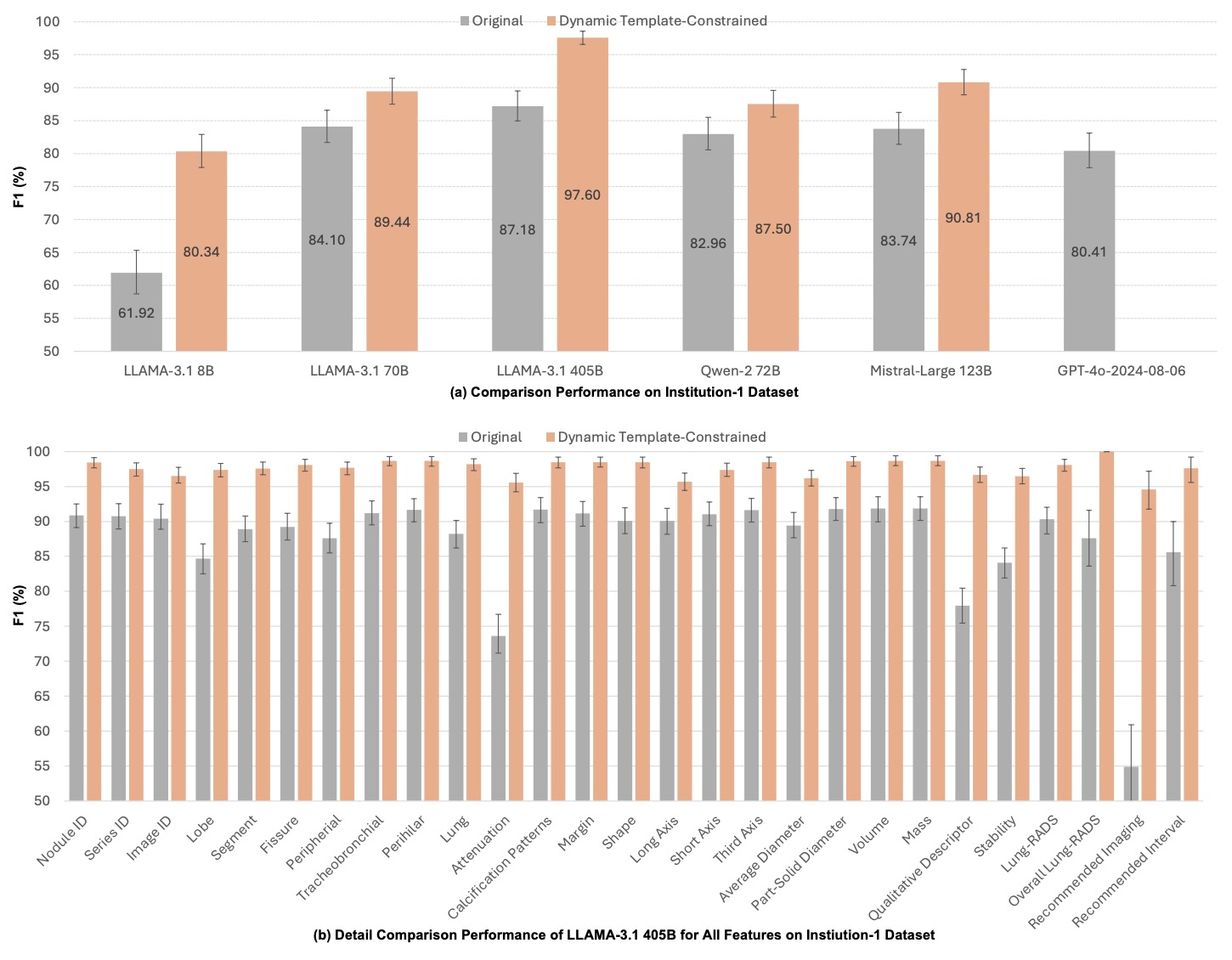
Figure 6: Institution-1 performance comparison.
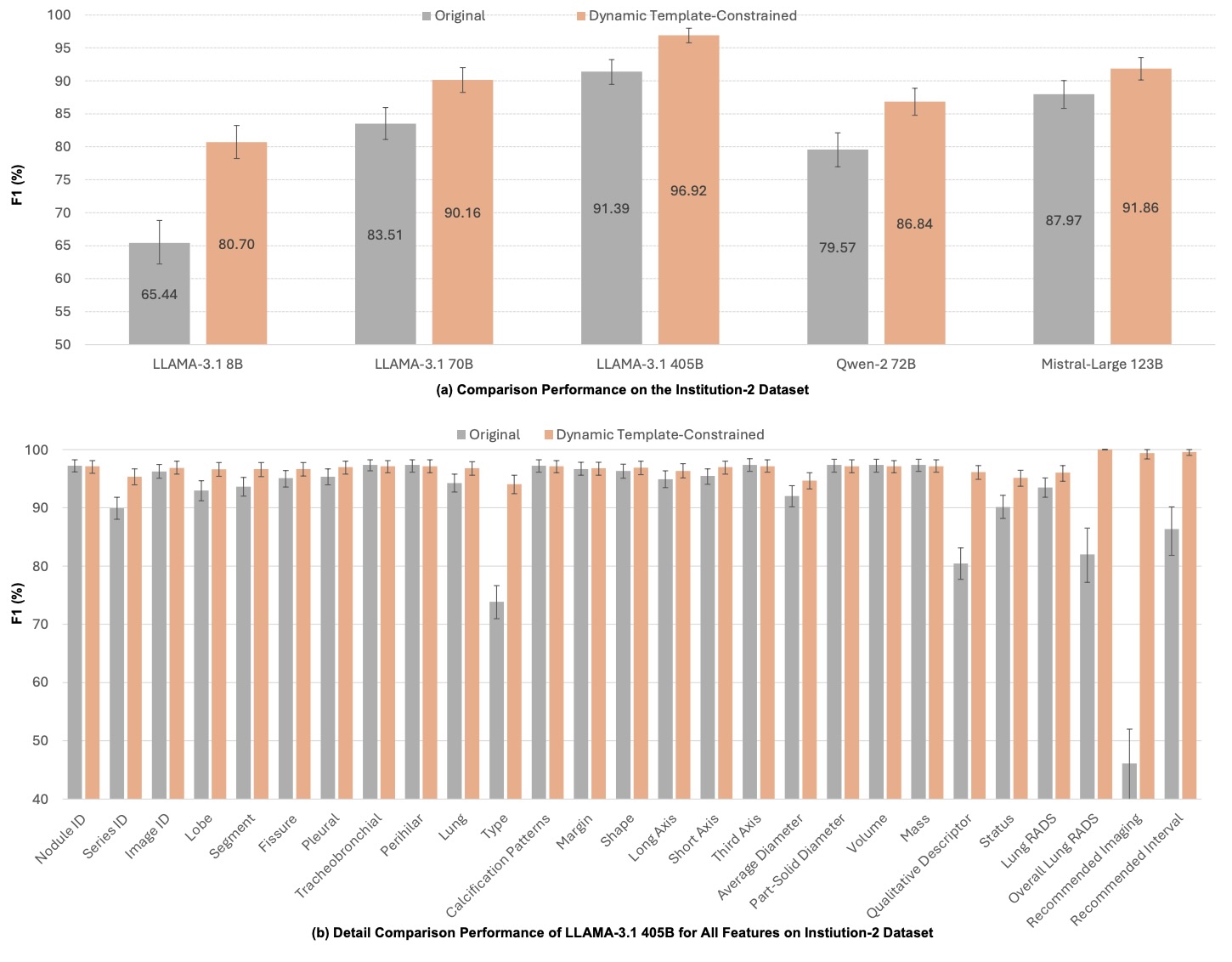
Figure 7: Institution-2 performance comparison.
📈 Extended Large-Scale Evaluation (Unpublished Update)
Beyond the cross-institutional validation reported in the manuscript, we further evaluated updated model variants on a 5,000-report dataset derived from our newly curated structured radiology corpus.
- LLaMA-3.1–8B (Template-Constrained): 98.05% F1 on 5k evaluation set
- LLaMA-3.2–1B (Fine-Tuned & Constrained): 94.27% F1 on 5k evaluation set
These results demonstrate that:
- The constrained decoding framework scales effectively to larger evaluation sets
- An 8B model achieves near-clinical-grade performance (>98% F1)
- Even a lightweight 1B model maintains strong structural fidelity (>94% F1)
- Model size-performance trade-offs enable deployment flexibility
Note: These extended results are not included in the current arXiv manuscript version but reflect subsequent internal evaluation.
🔎 Large-Scale Statistical Mining
5,192 consecutive LDCT reports (3 years) were automatically converted to FSR.
Automatically derived statistics matched prior clinical literature:
- Upper lobes had significantly more nodules (p < 0.01)
- Right lung > left lung (p < 0.01)
- Ground-glass nodules more common in females (p < 0.01)
- Lung-RADS distribution consistent with prior studies
🔍 Nodule-Level Retrieval System
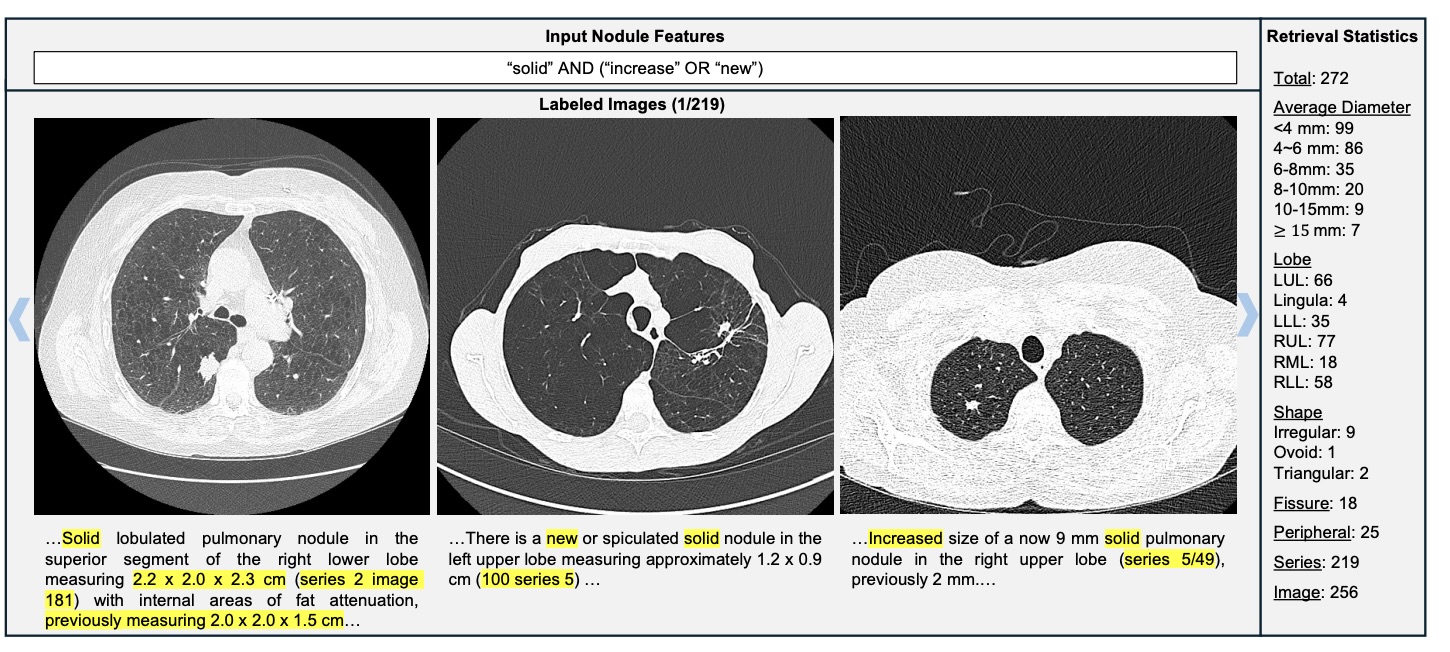
Figure 8: Complex semantic query: solid AND (increase OR new) → 272 nodules retrieved.
Unlike keyword matching, semantic reasoning detects growth even when "increase" is not explicitly mentioned.
⚙️ Synthetic Fine-Tuning & Efficient Model Scaling
To reduce computational cost and enable local deployment, a LLaMA-3.1–8B model was fine-tuned using a large-scale synthetic radiology dataset generated through template-driven structured sampling and free-text synthesis.
- Training samples: 189,950
- Testing samples: 49,965
- Mean IoU: 0.183 → 0.840
- Mean F1: 0.349 → 0.899
- Inference time (5,000 reports): 9h → 2.5h
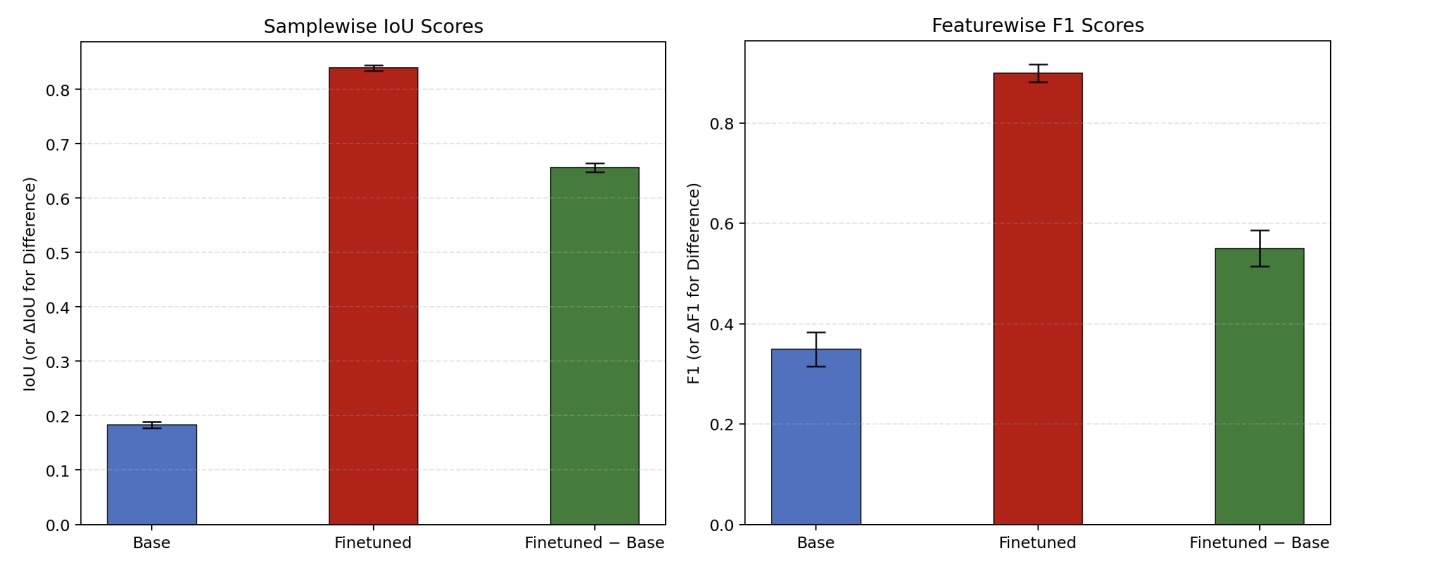
Figure 9: Structural IoU & feature-wise F1 improvement after fine-tuning.
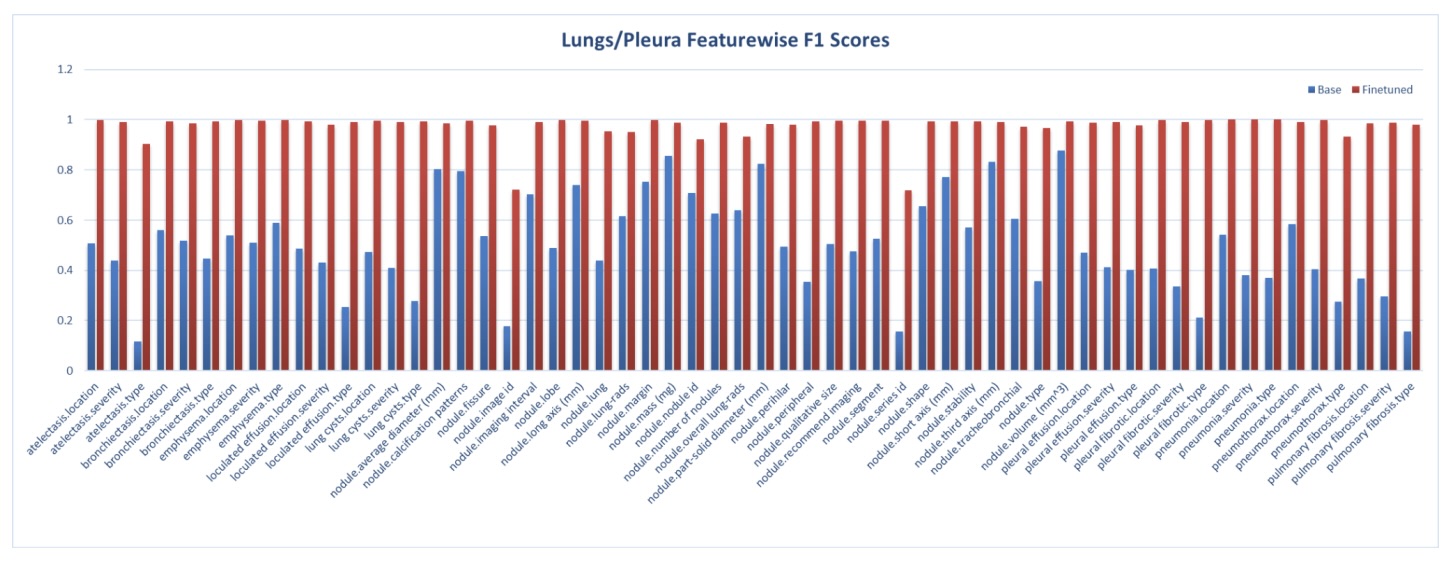
Figures 10: Lung/Pleura feature-level F1 improvements across anatomical systems.
🚀 Key Contributions
- Dynamic template-constrained decoding eliminating hallucinations
- Cross-institutional validation (n=7,442)
- Zero formatting errors
- Open-source vLLM-Structure framework
- Efficient fine-tuned 8B model for scalable deployment
- Automatic statistical mining & semantic retrieval