Hierarchical Attention Transformer (TrialDura)
Bio-BERT embeddings + hierarchical attention over eligibility criteria + regression head
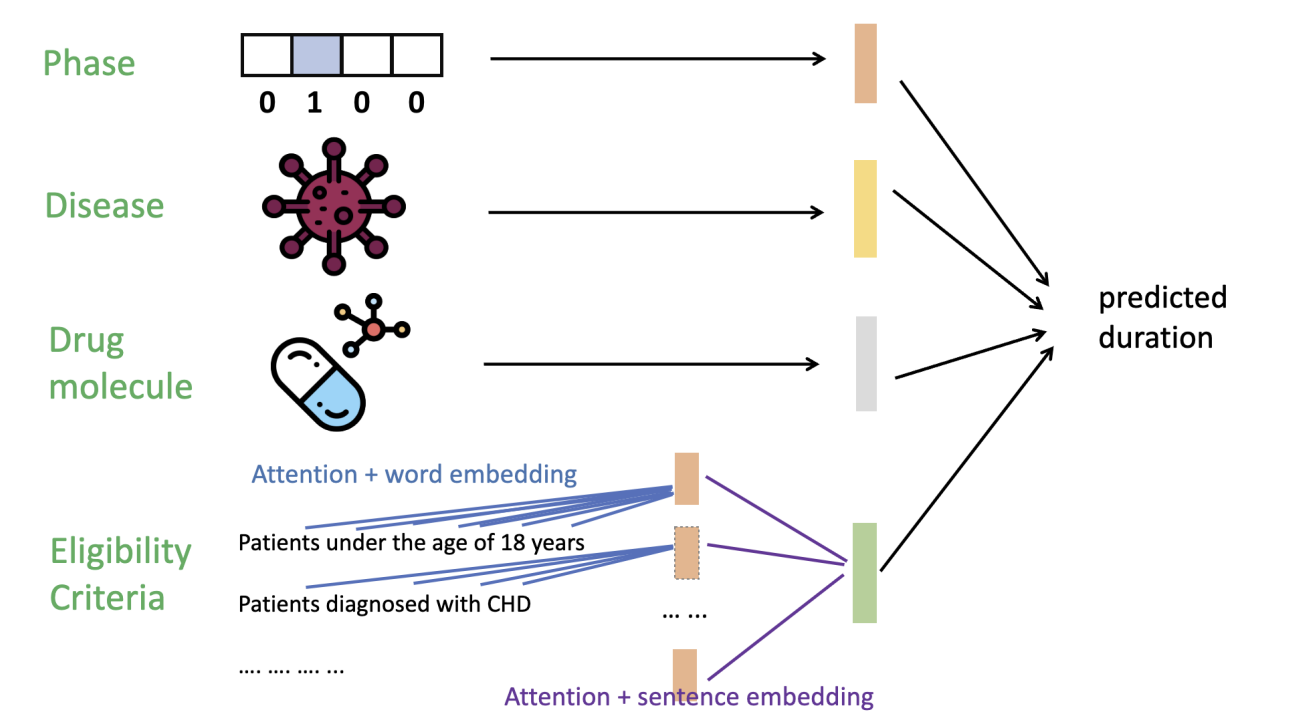

A hierarchical transformer architecture for modeling multi-level clinical trial features, enabling accurate and interpretable prediction of trial completion timelines.
NeurIPS 2024 Workshop on AI for New Drug Modalities
Interpretable, multimodal timeline forecasting for clinical trial planning and budgeting
Clinical trials are often long and expensive, so reliable duration prediction supports budgeting, staffing, recruitment planning, and operational risk management. TrialDura formulates duration prediction as a supervised regression problem and estimates trial duration (years) using multimodal trial records.
Inputs: (1) trial phase (I–IV, one-hot), (2) disease set (ICD-10 codes), (3) drug molecules (drug names), and (4) eligibility criteria (inclusion + exclusion text). TrialDura embeds drug/disease/criteria text with Bio-BERT and models criteria using hierarchical attention to obtain interpretable importance signals.
Dataset scale: 114,604 trials total, split temporally: training/validation trials starting before Jan 1, 2019 and test trials starting after. The paper reports 77,818 training records and 36,786 testing records.
Bio-BERT embeddings + hierarchical attention over eligibility criteria + regression head


Benchmarking against classical ML and neural baselines on 114,604 ClinicalTrials.gov records
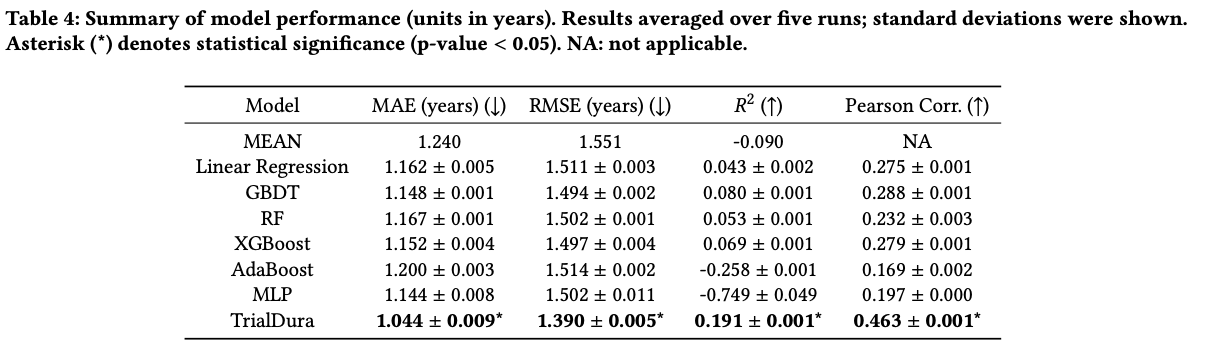
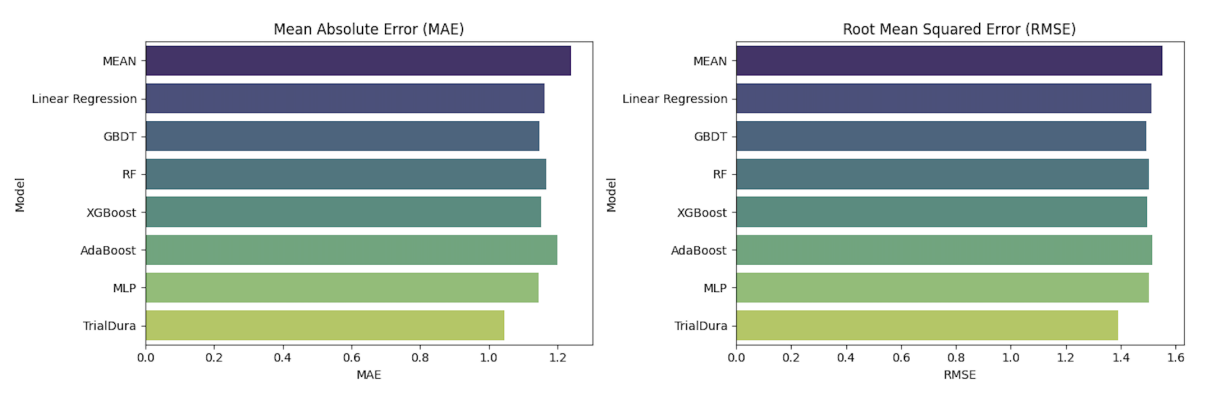
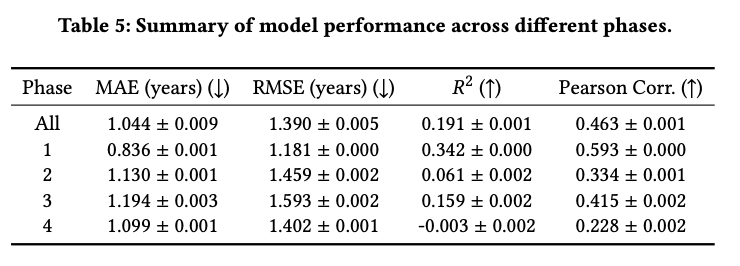
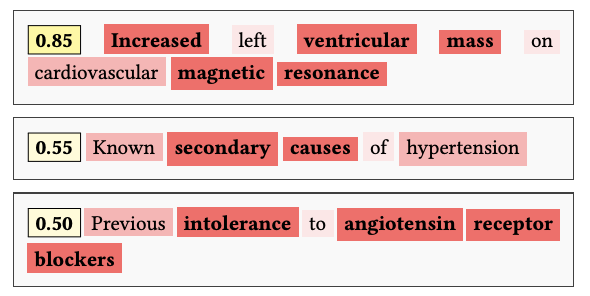
Identifying which eligibility criteria sentences/terms drive duration estimates

Understanding the contribution of unified training and sentence aggregation choices
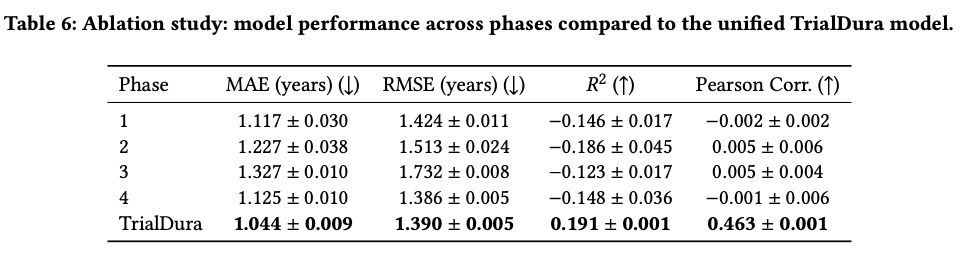
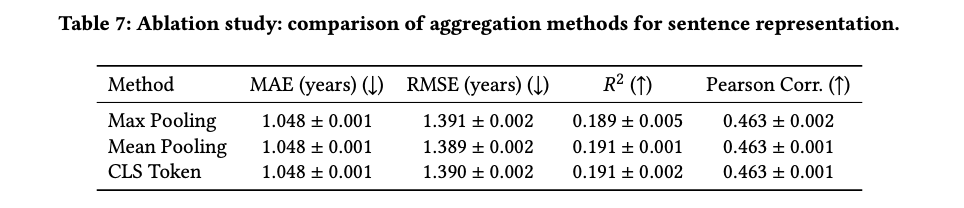
Examples across phases, diseases, and drugs
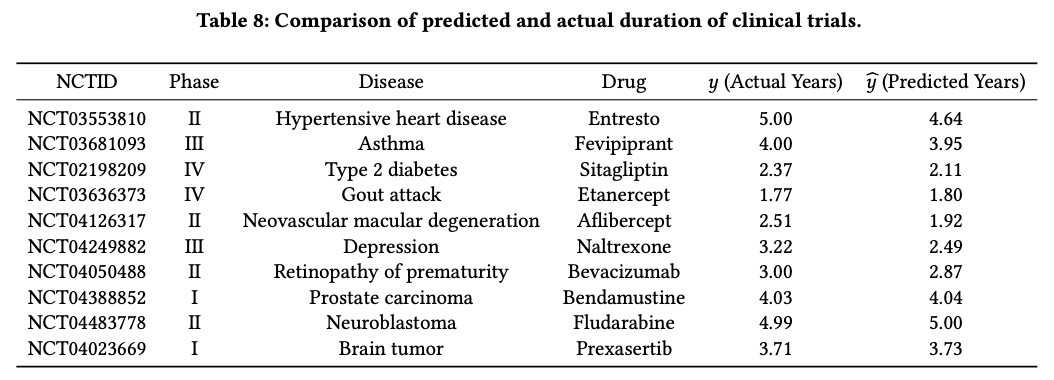
@article{yue2024trialdura,
title={TrialDura: Hierarchical Attention Transformer for Interpretable Clinical Trial Duration Prediction},
author={Yue, Ling and Li, Jonathan and Islam, Md Zabirul and Xia, Bolun and Fu, Tianfan and Chen, Jintai},
journal={arXiv preprint arXiv:2404.13235},
year={2024}
}